Harnessing the Power of Bioinformatics to Advance Research
Two bioinformaticists are dedicated to improving research by applying computer science, mathematics and statistics to analyze biological data.
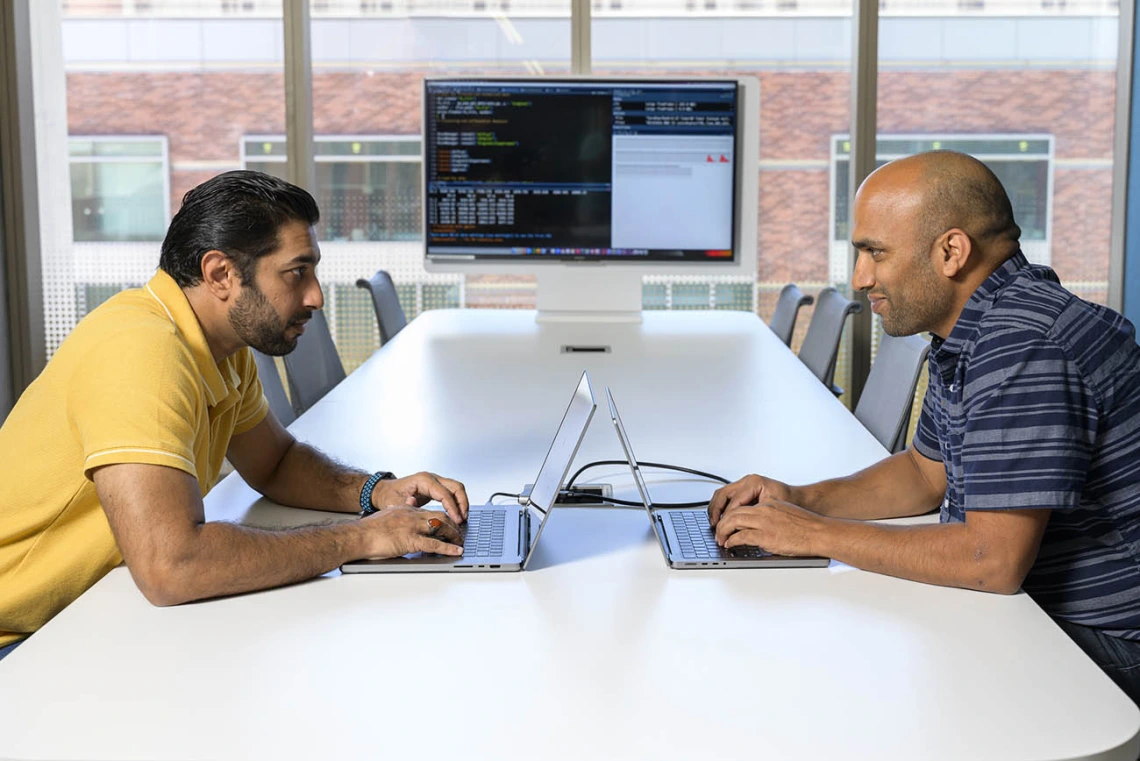

Syed Shujaat Ali Zaidi, PhD, (left) and Rudramani Pokhrel, PhD (right), are working with researchers to apply computational techniques to rapidly accelerate the pace of host-microbe research as part of Personalized Defense, a University of Arizona Health Sciences initiative.
Mathematics has long been used to explain, understand, and advance medical sciences. In the late 1400s, Italian polymath Leonardo da Vinci used ratios to accurately depict the human body in drawings, and calculus played an integral role in turning X-ray data into readable images of what is inside the body. In recent history, computer science and statistics made it possible to sequence and map all 3.2 billion letters of the human genome.

As a bioinformaticist, Syed Shujaat Ali Zaidi, PhD, combines technical skills with a passion for finding computational solutions to important biological questions.
At the University of Arizona Health Sciences, Syed Shujaat Ali Zaidi, PhD, and Rudramani Pokhrel, PhD, both bioinformaticists in the UArizona College of Medicine – Tucson’s Department of Immunobiology, are employing their skills as part of a strategic initiative to create defenses against disease.
“If we can find differences or variations between diseased versus normal population, it could be a great benefit for preventing the disease or developing the drug and therapeutics,” Dr. Pokhrel said.
“Identifying novel microbes from various habitats and using their products to treat diseases like diabetes, cancer, high cholesterol and many other neurological disorders would be extremely beneficial,” Dr. Zaidi added. “Metagenomics is a new way or dimension of understanding, analyzing and treating diseases.”
Their main objectives as part of the UArizona Health Sciences Personalized Defense strategic initiative are to advance and support research and to create best practices and workflows for analyzing data.
Advancing research across disciplines
Bioinformatics is an interdisciplinary powerhouse that has the potential to better research across fields.
“At some point, a researcher will need bioinformatics support to analyze biological data, which if done manually, is extremely time consuming and error prone,” Dr. Zaidi said. “It could be data integration, analysis or visualization, and comparative study.”

Rudramani Pokhrel, PhD, designs and implements bioinformatics analysis strategies for researchers, with a focus on using single-cell multi-omics and next-generation sequencing data to characterize immune responses, define existing microbes and discover novel infectious pathogens.
“The big question we have to answer from different perspectives is how bacteria and viruses are interacting with each other and within the host,” Dr. Zaidi said. “How are they causing or preventing disease?”
Some of the research Dr. Pokhrel is supporting is examining cytomegalovirus, a herpesvirus found in more than 50% of U.S adults over 40, according to the Centers for Disease Control and Prevention.
“We are looking at the cellular microenvironment – how the different kinds of cells are affected by this virus,” Dr. Pokhrel said. “Once we have the outcome from the data we can move forward, targeting those pathways or diseases.”
He also collaborates with researchers at the UArizona Cancer Center who are studying breast cancer using RNA sequencing data. Eventually, they hope to identify biomarkers that would allow them to diagnose breast cancer earlier and develop novel treatment opportunities.
Dr. Zaidi is working with Bonnie LaFleur, PhD, research professor, associate director of the Center for Biomedical Informatics and Biostatistics Data Resource Center and BIO5 Institute member, to identify how different treatments affect metabolites in the spinal cord and kidney.
“Changes in metabolite will cause alterations in pathways,” he said. “That is what we are trying to observe.”
Developing bioinformatics best practices
In addition to working with researchers, Drs. Zaidi and Pokhrel are tasked with determining and implementing bioinformatics best practices to ensure the research results can be replicated. Even the smallest deviation in workflow or process can skew results when dealing with such immense amounts of data.
“In any kind of data analysis field, reproducibility is a major challenge,” Dr. Zaidi said. “Every lab has its own techniques and way of doing work. What we are trying to do is develop a best practice for each of the data type.”

Drs. Zaidi and Pokhrel collaborate with bioinformatics and data science specialists across campus, including in the UArizona Cancer Center, Data Science Institute, CyVerse, Steele Childern’s Center Microbiome Reseach group and Center for Biomedical Informatics and Biostatistics.
T cells are a type of white blood cell found in the immune system. The T cell receptor allows a T cell to recognize different pathogens and tumors. Sequencing T cell receptors can be a powerful tool to examine the scope of the immune response to infection.
“This will help the researchers analyze the sequencing data,” Dr. Zaidi said, adding that the program also generates publication-ready visualizations for researchers to submit to scientific journals.
Drs. Zaidi and Pokhrel both say they are happy to be on the forefront of an exciting field that continues to grow. But for them, the most rewarding part is thinking about the practical applications of the research made possible through bioinformatics.
“It is a great satisfaction that I am taking some steps to make people’s lives better,” Dr. Pokhrel said.
Dr. Zaidi agreed and added, “My motivation is to explore nature and do something beneficial for mankind. I don’t want to just publish research; I want to come up with something that helps people improve their lives.”
Our Experts
Contact
Brian Brennan
702-604-3088
brianbrennan@arizona.edu

